Physiological Process and Mechanism of Senescence in Mutant Rice with Functional Deletion on OsVHA-A1 Gene
-
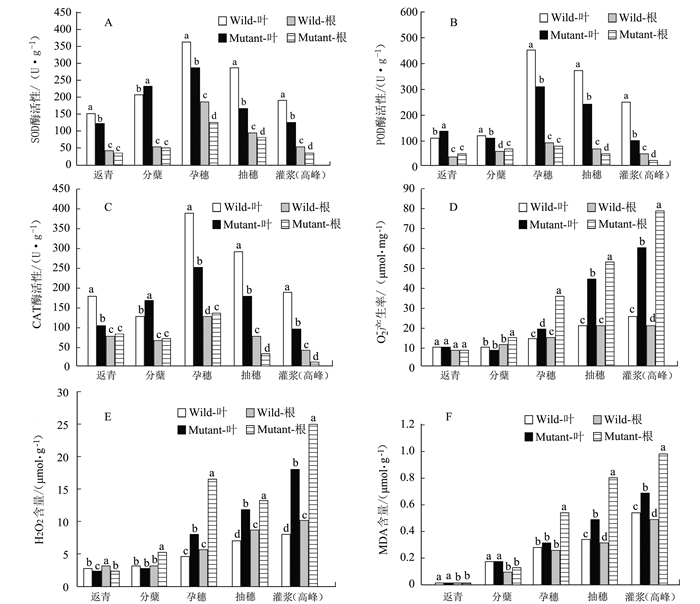
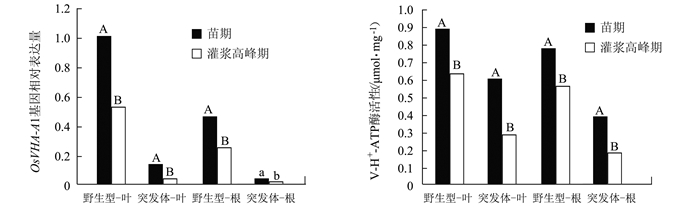
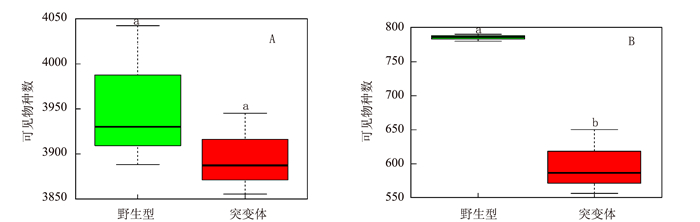
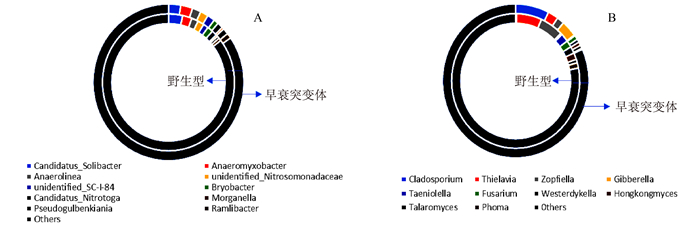
摘要: 负责编码水稻液泡膜ATP酶A亚基的OsVHA-A1基因在水稻生长发育和抵抗环境压力等诸多方面发挥重要作用。本试验以筛选纯合的OsVHA-A1基因缺失突变体株系及其野生型为材料,发现OsVHA-A1基因缺失突变会导致该突变体水稻生育后期发生严重的早衰现象与减产。两种基因型水稻叶片与根系中抗氧化保护酶活性随生育期呈现动态变化,孕穗期后突变体根系和叶片的衰老指标(包括H2O2含量、O2-产生率和MDA含量)均明显高于野生型;与野生型对比,突变体根系衰老程度显著高于其叶片。在早衰前后两个时期,即苗期和灌浆高峰期,对OsVHA-A1基因进行qRT-PCR定量,发现灌浆高峰期时的突变体与野生型叶片和根系中的OsVHA-A1基因相对于苗期均下调表达,V-ATPase酶活性也显著下降,突变体下降幅度大于野生型。利用16S扩增子分析根际土壤微生物组成时发现,灌浆高峰期突变体根际土壤中植物病原菌的含量显著上升。同时,HPLC技术测得突变体根际土壤中对羟基苯甲酸和对香豆酸两种酚酸类物质极显著低于野生型。说明不同基因型水稻根际土壤微生物存在对根系分泌物的选择性利用并诱导根际病原菌的增长。相关性分析结果表明,孕穗期后两种基因型水稻的根系活力下降变化与其功能叶片叶绿素含量及光合速率降低呈一定的相关关系。综上所述,OsVHA-A1基因缺失突变直接介导了水稻根系和叶片早衰的发生,并通过选择性诱导根际病原菌增长而加剧突变体根系与整体老化。Abstract: Vacuolar H+-ATPases (V-ATPase) is a multi-subunit enzyme complex found in the tonoplast of eukaryotes. V-ATPase subunitA(VHA) encoded by OsVHA-A1 gene is crucial in the development and tolerance to environmental stress of rice. This study applied a wild type and a purified mutant rice with a single cytosine deletion from its OsVHA-A1 gene to compare the differences of their phenotypes during the entire growth period. The mutant rice senesced at late-growth stage resulting a significant yield reduction as compared to the wild type. The activities of antioxidant enzymes including SOD, POD and CAT of the two isogene rice lines showed differential temporal patterns.The senescence-induced increases on O2- production as well as H2O2 and MDA contents in the leaves and roots of the mutant rice after booting stage were generally higher than those of the wild type.The performance of the roots tended to be poorer than that of the leaves in the mutant type in comparison with that of the wild type. OsVHA-A1 was significantly down-regulated and the V-ATPase activity significantly declined at grain-filling stage after senescence from seeding stage before senescence for both lines. But the down-regulated OsVHA-A1 expression and reduced V-ATPase activity in the roots and leaves were greater in the mutant rice than those in the wild type. It appeared that OsVHA-A1 played a crucial role in regulating rice senescence.Aside from genetic changes, environmental factors also contributed to the heightened root senescence, as the 16S rDNA sequencing showed that the pathogenic fungi population in the rhizosphere soil increased significantly when the mutant type at its grain-filling stage. In the soil, HPLC analysis identified 5 phenolic acids, i.e., phydroxybenzoic, vanillin, syringate, pcoumaric acid, and ferulic acid. The contents of phydroxybenzoic and pcoumaric acid were extremely significantly lower associated with the mutant than the wild type. It suggested that the pathogens might participate in the process of root senescence. Furthermore, the declined root activity was found correlating with the chlorophyll or photosynthesis decrease in the isogen rice lines after booting stage.It was concluded that the absence of functional OsVHA-A1 directly affected the senescence, whereas the altered underground microbial community further hastened the aging process of the rice plant.
-
Keywords:
- senescence /
- antioxidant enzymes /
- V-ATPase /
- rhizosphere microorganism /
- phenolic acid
-
0. 引言
【研究意义】猪是重要的经济畜禽之一,而脂肪沉积是影响猪肉质量和风味等的关键因素之一[1]。脂肪沉积不仅受到环境因素的影响,例如饲养方式、生长环境等,还与控制目标性状功能基因的表达和调控等遗传因素有着密切关系。此外,饲料营养水平的调节也可以影响猪的脂肪沉积。对于猪的脂肪沉积的研究不仅有利于提高猪肉质量和营养价值,还可以为猪的养殖提供理论依据和技术支持。有研究表明,脂肪代谢相关基因如ACACA和FASN等,它们表达量的差异对猪的脂肪沉积具有影响[2,3],包括影响猪肉中的饱和、不饱和脂肪酸含量。此外,研究表明,猪的品种、遗传背景和基因多态性等因素也会影响猪的脂肪沉积性状[4,5]。【前人研究进展】脂肪酸结合蛋白3(Fat acid binding proteins 3,FABP3),又称为心脏型脂肪酸结合蛋白[6],定位于猪第6号染色体区域,相对分子质量为15 kDa,主要参与脂肪酸的转运和利用等,FABP3是编码肌肉细胞内脂肪酸结合蛋白的基因,在畜禽的脂肪代谢中发挥着重要作用[7]。研究表明FABP3基因多态性与猪脂肪性状具有相关性,与猪肌内脂肪(Intramuscular fat,IMF)含量呈正相关[8,9]。有研究发现FABP3基因的表达水平与IMF含量同样显著相关[10]。LI等[11]通过分析猪体外成脂分化过程中FABP3基因的表达水平,发现该基因表达水平在分化过程中呈上调的趋势,表明FABP3可能在脂肪细胞功能中发挥重要作用。FABP3基因是影响猪肌内脂肪含量的候选基因[12]。研究表明,FABP3基因参与前脂肪细胞分化和脂肪沉积等过程,对提高棕色脂肪组织的耐寒性和脂肪酸氧化非常重要[13]。张敏[14]对FABP3在奶牛乳腺上皮细胞脂质合成中的研究表明,FABP3通过调节固醇调节元件结合蛋白1(SREBP1)、过氧化物酶体增殖物激活受体(PPARG)基因的mRNA及蛋白表达水平控制脂滴的形成;JIANG等[15]同样发现miR-3-4789p可通过抑制FABP3基因表达来抑制前脂肪细胞分化和脂滴沉积;CHMURZYNSKA等[16]、BLECHA等[17]、LI等[18]、WANG等[19]通过挖掘FABP3基因的多态性位点发现该基因与脂肪性状相关。【本研究切入点】藏猪是我国唯一的高原、高寒放牧猪种,具有耐粗饲、耐寒以及沉脂能力强等生物学特性[20,21]。这些特征使得藏猪的脂肪沉积方式可能与低海拔生态区域的猪不同,研究藏猪的脂肪沉积可以为高海拔地区的畜牧业发展提供科学依据。FABP3基因在脂肪沉积过程中发挥重要作用,对功能基因的深入了解对提高畜禽肉品质具有重要价值,目前关于FABP3基因的多态性和表达水平对藏猪肉品质的影响鲜有报道。【拟解决的关键问题】为加快对藏猪肉品质改良进程,本研究以饲养在西藏林芝(平均海拔2900 m)的藏猪和大约克猪为研究对象,采用一代测序技术对FABP3基因的5'侧翼区和CDS区进行测序和多态性分析,并利用RT-qPCR技术,比较FABP3基因在藏猪和大约克猪肝脏、背最长肌以及背脂中的表达情况,以期为猪FABP3基因调控脂肪代谢的分子机制奠定基础。
1. 材料与方法
1.1 试验材料
采集西藏农牧学院教学实习牧场180日龄的藏猪(n=28)和大约克猪(n=39)耳组织用于进行DNA提取。挑选无血缘关系的藏猪和大约克猪各10头同期饲养至180日龄进行屠宰,取其肝脏、背脂和背最长肌组织置于注有RNA保存液的样品管中,−80 ℃保存,用于RNA提取。
1.2 组织DNA、RNA提取及cDNA的制备
采用苯酚-氯仿抽提法提取DNA[22],使用Trizol法提取藏猪与大约克猪组织总RNA[23];用NanoDrop One超微量分光光度计检测所提DNA和RNA的纯度和浓度;1%琼脂糖凝胶电泳检测RNA的完整性。后采用FastKing cDNA第一链合成试剂盒(天根)制备cDNA,−20 ℃保存,用于后续荧光定量试验。
1.3 DNA引物设计与合成
登录NCBI(htttp://www.ncbi.nlm.nih.gov)网站下载猪FABP3基因(登录号:NC_010448.4)起始密码子上游3000 bp区域DNA序列与CDS区(登录号NM_001099931.1)序列,根据序列信息进行引物设计(Primer 5),并由上海生工生物工程有限公司进行引物合成,引物信息如表1所示。
表 1 FABP3基因5'侧翼区和CDS区引物序列Table 1. Primer sequence of 5' flanking region and CDS region in FABP3引物
Primer扩增区域
Amplified region引物序列(5′-3′)
Primer sequences退火温度
Annealing temperature/℃产物大小
Product length/bp5′−FABP3-1 356 bp至−460 bp F: TCAGCCCAAGAGTGAGTTTC
R: CCTTCTTCCTCGAAAGCG56 817 5′−FABP3-2 −430 bp至−1365 bp F: TCTGCTGGCTCAAGTTCAGT
R: GAGAGGAGAAAGGAAACTCACT58 953 5′−FABP3-3 −1342 bp至−2197 bp F: TAGGAGTCAACTTTGGTGAGC
R: CCAACTGAACTTGAGCCAGCA59 856 5′−FABP3-4 −2195 bp至−3033 bp F: CTGGGAACCTCCATATGTCG
R: CTAAGCCACAATCTATCACCT57 849 FABP3-CDS 34 bp至445 bp F:CCTGTTCTGTCGTCTCTTTCTCA
R:TGCCTCTTTCTCGTAAGTGCG60 440 1.4 荧光定量PCR引物设计与合成
在NCBI(htttp://www.ncbi.nlm.nih.gov/)下载猪FABP3基因(登录号NM_001099931.1)mRNA序列,选取GAPDH作为内参基因,利用软件Primer Premier 5.0设计引物(引物信息见表2),用于组织表达分析,并由上海生工生物工程有限公司进行合成。
表 2 FABP3基因定量PCR引物序列Table 2. Primer sequence of FABP3 for quantitative PCR基因名称
Gene name登录号
GeneBank number引物序列(5′-3′)
Primer sequences退火温度
Annealing temperature/℃产物大小
Product length/bpFABP3 NM_001099931.1 F: ATGACCAAGCCTACCACAA
R: AAGTTTGCCTCCATCCAGT57 171 GAPDH NM_001206359.1 F: CACCATCTTCCAGGAGCGAG
R: CCCTTCAAGTGAGCCCCG57 120 1.5 SNPs筛选与转录因子预测
对FABP3基因起始密码子上游3000 bp区域的DNA序列进行PCR扩增,经琼脂糖凝胶电泳检测后选取藏猪和大约克猪各10个合格的PCR产物进行混池测序,利用Chromas Pro软件对测序结果进行分析、筛选SNPs位点。针对筛选的SNPs进一步送扩大个体再次进行测序。综合两次测序结果计算各突变位点的基因型频率与基因频率。登录JASPAR(htttp://jaspar.binf.ku.dk/)转录因子预测网站进行突变位点转录因子预测。
1.6 荧光定量PCR
以cDNA为模板,选取藏猪与大约克猪各10个个体进行RT-qPCR,每个个体设置3个重复,反应体系为20 μL。并使用2−△△ct法计算FABP3基因在组织中的相对表达量。
1.7 统计分析
使用SPSS 26.0软件对FABP3基因mRNA相对表达量进行单因素方差分析,测定结果以P值(P value)表示;对FABP3基因的基因型频率和基因频率进行卡方检验。P <0.05为差异显著,P < 0.01为差异极显著。
2. 结果与分析
2.1 藏猪与大约克猪组织DNA检测
从琼脂糖凝胶电泳检测结果(图1)可以看出,所提DNA样品条带比较清晰完整,可以用于后续扩增试验。
2.2 FABP3基因SNPs筛选
对测序结果分析发现FABP3基因在5’侧翼区共有2个突变位点C-635A和T-114C(图2)。突变位点C-635A在藏猪中存在AA基因型,在大约克猪中存在CC、CA和AA基因型,等位基因A为优势基因。突变位点T-114C在藏猪中存在TC、CC基因型,在大约克猪中存在TT和TC基因型,等位基因C为优势基因(表3)。藏猪和大约克猪FABP3基因CDS区均未发现突变。
表 3 FABP3基因SNPs位点基因型频率及卡方检验Table 3. Genotype frequency and chi-square test of SNPs on FABP3位点
Loci品种
Species样本量
Sample size基因型频率(个体数/频率)
Genotype frequency (Individuals/Frequency)基因频率
Gene frequencyχ2值
CardinalityP值
P valueCC CA AA C A C-635A 大约克猪 39 16/0.410 16/0.410 7/0.179 0.615 0.385 0.693 0.707 藏猪 28 15/0.520 12/0.410 28/1.000 0.000 1.000 0.000 1.000 藏猪 vs 大约克猪 χ2=43.979;P<0.01 T-114C TT TC CC T C 大约克猪 39 33/0.846 6/0.154 0/0 0.923 0.077 0.271 0.873 藏猪 28 0/0.000 1/0.036 27/0.964 0.018 0.982 0.009 0.995 藏猪 vs 大约克猪 χ2=63.476;P<0.01 2.3 FABP3基因的基因频率与基因型频率
由表3可知,FABP3基因的突变位点在藏猪和大约克猪种内均符合哈迪-温伯格平衡定律(P>0.05)。藏猪和大约克猪在C-635A、T-114C位点呈极显著差异(P<0.01)。
2.4 转录因子预测
如表4所示,通过对转录因子预测发现SNPs位点碱基突变前后有部分转录因子结合位点消失和新转录因子产生。FABP3基因起始密码子上游3000 bp区域中存在2个SNPs位点(C-635A和T-114C),其中C-635A位点上C向A的突变导致ARNT2、SUMO2、 RB1、 CTCF、SMAD4、LF11、CHD1、SUZ12、YY1结合位点消失,并出现了新的转录因子STAT5B、TCF7L2、TCF12、SREBP1、MYH11、SPI1、TP53、TBX21、HOXA9;T-114C位点上T向C的突变导致E4F1结合位点消失,并出现了新的转录因子HES5、CLOCK、WT1、EP300、THAP11、KLF5。
表 4 FABP3基因SNPs位点转录因子预测结果Table 4. Predicted transcription factors in FABP3 SNPs突变位点
Mutation Loci突变前序列
Pre-mutation sequence突变后序列
Post-mutation sequence消失转录因子
Disappearance of transcription factors新增转录因子
Addition of transcription factorsC-635A TGGGGCGGGGG TGGGGAGGGGG ARNT2、 SUMO2、 RB1、 CTCF、SMAD4、LF11、CHD1、SUZ12、YY1 STAT5B、TCF7L2、TCF12、SREBP1、MYH11、SPI1、TP53、TBX21、HOXA9 T-114C ACGCCTCGTCA ACGCCCCGTCA E4F1 HES5、CLOCK、WT1、EP300、THAP11、KLF5 2.5 藏猪与大约克猪组织RNA电泳检测
从图3可以看出,RNA电泳图中28S、18S和5S条带清晰完整,表明本次提取的RNA完整无降解,满足后续反转录试验要求。
![]() 图 3 组织总RNA琼脂糖凝胶电泳M:Marker;1~3分别为藏猪肝脏、背脂、背最长肌组织;4~6分别为大约克猪肝脏、背脂、背最长肌组织。Figure 3. Agarose gel electrophoresis of total RNA in tissueM:Marker; 1–3: Liver, dorsal fat, and longest dorsal muscle tissues of Tibetan pig, respectively; 4–6: liver, dorsal fat, and longest dorsal muscle tissues of Yorkshire pig, respectively.
图 3 组织总RNA琼脂糖凝胶电泳M:Marker;1~3分别为藏猪肝脏、背脂、背最长肌组织;4~6分别为大约克猪肝脏、背脂、背最长肌组织。Figure 3. Agarose gel electrophoresis of total RNA in tissueM:Marker; 1–3: Liver, dorsal fat, and longest dorsal muscle tissues of Tibetan pig, respectively; 4–6: liver, dorsal fat, and longest dorsal muscle tissues of Yorkshire pig, respectively.2.6 FABP3基因在不同组织中的mRNA表达量
利用RT-qPCR技术分别对FABP3基因在藏猪和大约克猪两猪种肝脏、背脂和背最长肌3个组织中的表达水平进行检测(图4)。FABP3基因在藏猪的肝脏和背最长肌组织中的mRNA表达水平极显著高于大约克猪(P < 0.01),在背脂组织中的mRNA表达水平显著高于大约克猪(P < 0.05)。
![]() 图 4 FABP3基因在藏猪、大约克猪肝脏、背脂和背最长肌中的mRNA相对表达量TP为藏猪,YY为大约克猪;*为显著差异(P < 0.05),**为极显著差异(P < 0.01)。Figure 4. Relative expressions of FABP3 in liver, back fat, and longissimus dorsi muscle of TP and YYTP: Tibetan pig; YY: Yorkshire pig; *: significant difference at P<0.05;**: extremely significant difference at P<0.01.
图 4 FABP3基因在藏猪、大约克猪肝脏、背脂和背最长肌中的mRNA相对表达量TP为藏猪,YY为大约克猪;*为显著差异(P < 0.05),**为极显著差异(P < 0.01)。Figure 4. Relative expressions of FABP3 in liver, back fat, and longissimus dorsi muscle of TP and YYTP: Tibetan pig; YY: Yorkshire pig; *: significant difference at P<0.05;**: extremely significant difference at P<0.01.3. 讨论与结论
本研究在FABP3基因起始密码子上游3 kb区域发现2个SNPs位点(C-635A和T-114C),在藏猪和大约克猪中,这两个SNP位点的等位基因频率和基因型频率差异极显著(P < 0.01)。后续对2个位点突变前后转录因子进行预测,发现在C-635A位点,当C突变为A后,转录因子CTCF消失,CTCF(CCCTC位点结合蛋白)在调控动物生长发育过程中发挥重要作用[24,25]。肖成等[26]在对小寒尾羊的研究中发现S100钙结合蛋白A1(S100 Calcium Binding Protein A1,S100A1)的表达量随着脂肪组织中新细胞的增大而出现显著升高,而CTCF可以抑制S100A1的活性[27]。因此,推测CTCF对脂肪的生成可能起到负调控的作用。当C突变为A后,在该位点结合产生新的转录因子——胆固醇调节元件结合蛋白1(SREBP1)。SREBP1是脂肪合成酶的主要调节剂[28],是控制脂肪合成的关键转录因子,降低SPRBP1的表达可以降低脂肪沉积[29,30]。在T-114C位点,当T突变为C,转录因子ZnF蛋白E4F转录因子1(E4F1)消失。E4F1是早期胚胎发育过程中必需的多功能蛋白,在不同细胞类型增殖中发挥重要作用[31],包括干细胞。该转录因子可以与PARP-1基因共同作用来减少前体脂肪细胞的形成[32]。当T突变为C后,在该位点结合产生新的转录因子KLF5。CERVANTE等[33]在对山羊前体脂肪细胞的诱导试验中发现,KLF5在前体脂肪细胞的诱导分化中发挥着正向调控作用。KLF5(锌指蛋白转录因子5)可能调控脂肪细胞的生长和分化[34,35]。综上所述,该位点突变前后转录因子的变化可能对猪脂肪沉积产生影响 。
SCHAAP等[36]对小鼠的研究发现,敲除FABP3会降低小鼠对脂肪酸的吸收,使得小鼠体内脂肪量减少。YI 等[37]研究发现FABP3的过表达可以促进莱芜猪前脂肪细胞增殖;CHO K H等[38]通过对FABP3基因与大约克猪肌内脂肪含量的相关性研究发现,该基因与猪肌内脂肪含量呈正相关。本试验采用RT-qPCR技术分别对藏猪和大约克猪的背脂、肝脏和背最长肌3个组织中FABP3基因的表达水平进行检测,发现在肝脏和背最长肌组织上藏猪FABP3基因的mRNA相对表达量均极显著高于大约克猪;在背脂组织中,藏猪FABP3基因的mRNA相对表达量显著高于大约克猪。FABP3基因在藏猪中呈现高表达的趋势,而藏猪是典型的脂肪沉积型猪种,肌内脂肪含量较高,推测FABP3基因的高表达是藏猪脂肪能力强以及肌内脂肪含量高的原因之一,同时推测FABP3基因对猪的脂肪沉积有一定正向调控作用。
综上,藏猪FABP3基因5'侧翼区存在的2个突变位点C-635A和T-114C可能是参与脂肪生成和沉积的重要调控位点,FABP3基因的高表达可能正向调控藏猪脂肪沉积并对其肉品质产生重要影响。研究结果为后续阐明FABP3基因在猪脂肪沉积过程中的分子作用机制与分子标记辅助育种提供新思路和理论依据。
-
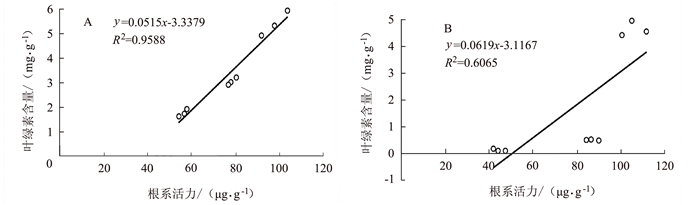
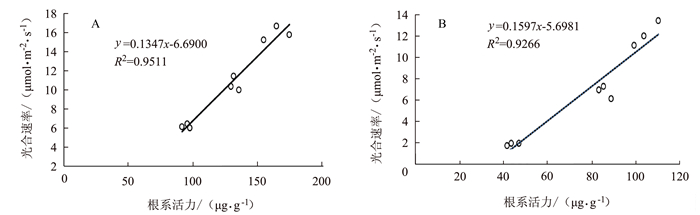
图 9 早衰后(孕穗期,抽穗期,灌浆高峰期)根系活力与叶绿素相关性分析
注:A为野生型,B为OsVHA-A1基因缺失突变体。图 10同。
Figure 9. Correlation between root activity and leaf chlorophyll content after senescence in different stages
表 1 OsVHA-A1基因缺失突变体与其野生型水稻农艺性状
Table 1 Agronomic traits of OsVHA-A1 mutant and wild type rice
水稻
类型株高
/cm根长
/cm地上部
/cm根干重
/g叶干重
/g有效穗
/穗结实率
/%千粒重
/g野生 127.3±2.5A 18.9±1.4a 108.5±2.1A 1.34±0.1A 4.75±1.5A 11.2±2.1A 88.3±4.6A 25.3±1.5A 突变 86.8±3B 15.8±1.2b 67.5±2.5B 0.73±0.1B 1.26±1.2B 5.8±1.5B 37.6±5.6B 13.5±1.4B 注:同列数据后不同大、小写字母分别表示差异达极显著(P<0.01)和显著水平(P<0.05)。 -
[1] 王复标, 黄福灯, 程方民, 等.水稻生育后期叶片早衰突变体的光合特性与叶绿体超微结构观察[J].作物学报, 2012, 38(5):871-879. http://d.old.wanfangdata.com.cn/Periodical/zuowxb201205014 [2] 白海齐.水稻灌浆期叶片衰老期间miRNA高通量测序及生物信息学分析[D].杭州: 杭州师范大学, 2015. [3] 郭士伟, 夏士健, 赵学强, 等.超级杂交稻两优培九及其亲本生育后期早衰的内源激素和营养生理研究[J].农业科学与技术:英文版, 2014(11):1914-1918. DOI: 10.3969/j.issn.1009-4229.2014.11.021 [4] 谢金水.生育后期养分胁迫对水稻衰老进程影响的蛋白质组学研究[D].南昌: 江西农业大学, 2012. [5] LIM P O, KIM H J, NAM HG. Leaf senescence[J]. Annu Rev Plant Biol, 2007, 58:115-136. DOI: 10.1146/annurev.arplant.57.032905.105316
[6] YOSHIDA S. Molecular regulation of leaf senescence[J]. Curr Opin Plant Biol, 2003(6):79-84. http://d.old.wanfangdata.com.cn/Periodical/yyxb201606007
[7] WANG P, SUN X, CHANG C, et al. Delay in leaf senescence of Malus hupehensis, by long-term melatonin application is associated with its regulation of metabolic status and protein degradation[J]. Journal of Pineal Research, 2013, 55(4):424-434. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=110e6484f23d5d5b50e5c840c4240890
[8] OZGUR R, UZILDAY B, SEKMEN A H, et al. Reactive oxygen species regulation and antioxidant drfence in halophytes[J]. Functional Plant Biology Fpg, 2013, 40(8-9):832-847. https://www.ncbi.nlm.nih.gov/pubmed/9328571
[9] HUA C, WANG R. Changes of SOD and CAT activities and MDA content during senescence of hybrid rice and three lines leaves[J]. Acta Botanica Boreali-occidentalia Sinica, 2003, 23(3):406-409. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=xbzwxb200303008
[10] LI Z, WANG F, LEI B, et al. Genotypic-dependent alteration in transcriptional expression of various CAT isoenzyme genes in esl, mutant rice and its relation to H2O2-induced leaf senescence[J]. Plant Growth Regulation, 2014, 73(3):237-248. DOI: 10.1007/s10725-013-9884-6
[11] UEDA Y, UEHARA N, SASAKI H, et al. Impacts of acute ozone stress on superoxide dismutase (SOD) expression and reactive oxygen species (ROS) formation in rice leaves[J]. Plant Physiology & Biochemistry Ppb, 2013, 70(1):396-402. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=156d9ed6b39060adf4dc146b3c58042f
[12] RIBEIRO C W, KORBES A P, GARIGHAN J A, et al. Rice peroxisomal ascorbate peroxidase knockdown affects ROS signaling and triggers early leaf senescence[J]. Plant Science, 2017, 263:55-65. DOI: 10.1016/j.plantsci.2017.07.009
[13] KIM J, CHANG C, TUCKER M L. To grow old:regulatory role of ethylene and jasmonic acid in senescence[J]. Frontiers in Plant Science, 2015, 6(20):20. http://www.ncbi.nlm.nih.gov/pubmed/25688252
[14] MAO J J, ZHAO C C, HUANG F D, et al. Physiological Characterization and Gene Fine Mapping of a Leaf Early Senescence and Salt-sensitive Mutant osles in Rice[J]. Acta Agronomica Sinica, 2014, 40(5):769-778. DOI: 10.3724/SP.J.1006.2014.00769
[15] HÄFFNER E, KONIETZKI S, DIEDERICHSEN E. Keeping Control:The Role of Senescence and Development in Plant Pathogenesis and Defense[J]. Plants, 2015, 4(3):449-488. DOI: 10.3390/plants4030449
[16] WANG F, LIU J, ZHOU L, et al. Senescence-specific change in ROS scavenging enzyme activities and regulation of various SOD isozymes to ROS levels in psf mutant rice leaves[J]. Plant Physiology & Biochemistry Ppb, 2016, 109:248-261. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=6e8e2f8b721ec0aab79aeb8825b13a86
[17] ZHANG J, ZHANG N, LIU Y X, et al. Root microbiota shift in rice correlates with resident time in the field and developmental stage[J]. Science China Life Sciences, 2018, 61(6):1-9. DOI: 10.1007/s11427-018-9284-4
[18] BERENDSEN R L, PIETERSE C M, BAKKER P A. The rhizosphere microbiome and plant health[J]. Trends in Plant Science, 2012, 17(8):478-486. DOI: 10.1016/j.tplants.2012.04.001
[19] EAST R. Microbiome:Soil science comes to life[J]. Nature, 2013, 501(7468):18-19. DOI: 10.1038/501S18a
[20] KUNDU P, BLACHER E, ELINAV E, et al. Our gut microbiome:the evolving inner self[J]. Cell, 2017, 171(7):1481-1493. DOI: 10.1016/j.cell.2017.11.024
[21] CLAESSON M J, JEFFERY I B, CONDE S, et al. Gut microbiota composition correlates with diet and health in the elderly[J]. Nature, 2012, 488(7410):178-184. DOI: 10.1038/nature11319
[22] 张惠莹. OsVHA-A基因生化及生物学功能研究与分析[D].重庆: 重庆大学, 2013. [23] FORSTER C, KANE P M. Cytosolic Ca2+ homeostasis is a constitutive function of the V-ATPase in Saccharomyces cerevisiae[J]. Journal of Biological Chemistry, 2000, 275(49):38245-38253. DOI: 10.1074/jbc.M006650200
[24] SHAO E, NISHI T, KAWASAKINISHI S, et al. Mutational analysis of the non-homologous region of subunit A of the yeast V-ATPase[J]. Journal of Biological Chemistry, 2003, 278(15):12985-12991. DOI: 10.1074/jbc.M212096200
[25] SHAO E, FORGAC M. Involvement of the nonhomologous region of subunit A of the yeast V-ATPase in coupling and in vivo dissociation[J]. Journal of Biological Chemistry, 2004, 279(47):48663-48670. DOI: 10.1074/jbc.M408278200
[26] YANG X, GONG P, LI K, et al. A single cytosine deletion in theOsPLS1gene encoding vacuolar-type H+-ATPase subunit A1 leads to premature leaf senescence and seed dormancy in rice[J]. Journal of Experimental Botany, 2016, 67(9):2761-2776. DOI: 10.1093/jxb/erw109
[27] BEAUCHAMP C, FRIDOVICH I. Superoxide dismutase. Improved assays and an assay applicable to acrylamide gel[J]. Anal Biochem, 1971, 44:276-287. DOI: 10.1016/0003-2697(71)90370-8
[28] ZHOU W, ZHAO D, LIN X. Effects of waterlogging on nitrogen accumulation and alleviation of waterlogging damage by application of nitrogen fertilizer and mixtalol in winter rape (Brassica napes L.)[J]. Journal of Plant Growth Regulation, 1997, 16:47-53. DOI: 10.1007/PL00006974
[29] MUÑOZ-MUÑOZ J L, GARCÍA-MOLINA F, GARCÍA-RUIZ P A, et al. Enzymatic and chemical oxidation of trihydroxylated phenols[J]. Food Chemistry, 2009, 113(2):435-444. DOI: 10.1016/j.foodchem.2008.07.076
[30] QUINTANILLA-GUERRERO F, DUARTE-VAZQUEZ M, GARCIA-ALMENDAREZ B, et al. Polyethylene glycol improves phenol removal by immobilized turnip peroxidase[J]. Bioresource Technology, 2008, 99(18):8605-8611. DOI: 10.1016/j.biortech.2008.04.031
[31] AEBI H. Catalase in vitro[J]. Methods Enzymol, 1984, 105(105):121-126. http://d.old.wanfangdata.com.cn/OAPaper/oai_doaj-articles_1ba2d838d4c25e398eab33b9f6a6e4a4
[32] NAKANO Y, ASADA K. Hydrogen Peroxide is Scavenged by Ascorbate-specific Peroxidase in Spinach Chloroplasts[J]. Plant & Cell Physiology, 1981, 22(5):867-880. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=HighWire000002692968
[33] CHAKRABARTY D, DATTA S K. Micropropagation of gerbera:lipid peroxidation and antioxidant enzyme activities during acclimatization process[J]. Physiol Plant, 2008, (30):325-331. DOI: 10.1007/s11738-007-0125-3
[34] KUMAR G, KNOWLES N R. Changes in Lipid Peroxidation and Lipolytic and Free-Radical Scavenging Enzyme Activities during Aging and Sprouting of Potato (Solanum tuberosum) Seed-Tubers[J]. Plant Physiology, 1993, 102(1):115. DOI: 10.1104/pp.102.1.115
[35] KE D, SUN G, WANG Z. Effects of superoxide radicals on ACC synthase activity in chilling-stressed etiolated mungbean seedlings[J]. Plant Growth Regulation, 2007, 51(1):83-91. DOI: 10.1007/s10725-006-9150-2
[36] BRADFORD M, BRADFORD M M, BRADFORD N. A rapid and sensitive method of microgram quantities of protein utilizing the principle of protein-dye binding[J]. Analytical Biochemistry, 1976, 72(S 1-2):248-254. DOI: 10.1016-0003-2697(76)90527-3/
[37] SCHMITTGEN T D, LIVAK K J. Analyzing real-time PCR data by the comparative Crmethod[J]. Nat Protoc, 2008(3):1101-1108.
[38] 吴丹, 赵立, 庞文生, 等.太子参根际土壤酚酸类自毒物质的分析测定[J].中国民族民间医药, 2017, 26(24):32-34. http://d.old.wanfangdata.com.cn/Periodical/zgmzmjyyzz201724010 [39] EDGAR R C. UPARSE:highly accurate OTU sequences from microbial amplicon reads[J]. Nature Methods, 2013, 10(10):996-998. DOI: 10.1038/nmeth.2604
[40] WANG Q, GARRITY G M, TIEDJE J M, et al. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy[J]. Applied & Environmental Microbiology, 2007, 73(16):5261. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=6c141ed427613641a3970c1b595b045f
[41] DESANTIS T Z, HUGENHOLTZ P, LARSEN N, et al. Greengenes, a chimera-checked 16S rRNA gene database and workbench compatible with ARB[J]. Applied & Environmental Microbiology, 2006, 72(7):5069-5072. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=37aa57e221455e0469915ba972b04513
[42] SOGIN M L, MORRISON H G, HUBER J A, et al. Microbial diversity in the deep sea and the underexplored "rare biosphere"[J]. Proceedings of the National Academy of Sciences of the United States of America, 2006, 103(32):12115-12120. DOI: 10.1073/pnas.0605127103
[43] 石岩, 位东斌.土壤水分胁迫对小麦根系与旗叶衰老的影响[J].西北植物学报, 1998(2):196-201. DOI: 10.3321/j.issn:1000-4025.1998.02.008 [44] 邵彩虹, 李瑶, 钱银飞, 等.养分胁迫对威优916生育后期根系衰老影响的蛋白质组学分析[J].华北农学报, 2013, 28(2):12-19. DOI: 10.3969/j.issn.1000-7091.2013.02.003 [45] 王占武, 李晓芝, 刘彦利.根际微生物对冬小麦根系发育及产量性状的影响[J].华北农学报, 2000, 15(Z1):51-54. DOI: 10.3321/j.issn:1000-7091.2000.Z1.011 [46] LIN S, HUANGPU J J, CHEN T, et al. Allelopathic potential and identification of allelochemicals in Pseudostellariae heterophylla rhizosphere soil in different crop rotations[J]. Allelopathy Journal, 2014, 33(2):151-161. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=6656a99ec212f2bba0468183646f7708
[47] 胡海军, 吴亚男, 鄂洋, 等.设施园艺连作障碍研究进展[J].安徽农业科学, 2016, 44(5):49-51. DOI: 10.3969/j.issn.0517-6611.2016.05.019 [48] YUMIN, YUJUN-WO, CAOPEI-GEN, et al. Agrochemical Characteristics of Soil for Continuous Cropping Lily[J]. Chinese Journal of Soil Science, 2004, 35(3):377-379. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=trtb200403032
[49] RICE E L. Allelipathy-an overview[M].Chemically Mediated Interactions Between Plants and Other Organisms. Springer us, 1985:81-105.
[50] CHOU C H.The role of Allelopathy in Phytochemical Ecology[J].Phytochemical Ecology, 1989(9):19-37. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=26dfd3d16dec8f77246ed8164088b797
[51] 周军建.不同水稻根系分泌物对根际土壤微生物群落多样性影响的研究[D].福州: 福建农林大学, 2007. [52] 李进春.高产水稻后期长相及提高光合作用的有效措施[J].宁夏农林科技, 2013, 54(5):14-14. DOI: 10.3969/j.issn.1002-204X.2013.05.006 [53] 刘道宏.植物叶片的衰老[J].植物生理学报, 1983(2):16-21. http://d.old.wanfangdata.com.cn/Periodical/xbzwxb200106037 [54] 王复标, 黄福灯, 程方民, 等.水稻生育后期叶片早衰突变体的光合特性与叶绿体超微结构观察[J].作物学报, 2012, 38(5):871-879. http://d.old.wanfangdata.com.cn/Periodical/zuowxb201205014 [55] 李木英, 石庆华, 郑伟, 等.杂交稻后期叶片早衰特征及其与叶片N含量和根系活力关系初探[J].江西农业大学学报, 2008, 30(5):000757-765. http://d.old.wanfangdata.com.cn/Periodical/jxnydxxb200805001 [56] WANG Q, GARRITY G M, TIEDJE J M, et al. Naive Bayesian classifier for rapid assignment of rRNA sequences into the new bacterial taxonomy[J]. Applied & Environmental Microbiology, 2007, 73(16):5261. http://www.wanfangdata.com.cn/details/detail.do?_type=perio&id=6c141ed427613641a3970c1b595b045f



 下载:
下载: