Cloning and Expression of PsWRKY33 Promoter in Nane
-
摘要:目的 探讨油柰(Prunus salicina lindley)WRKY33
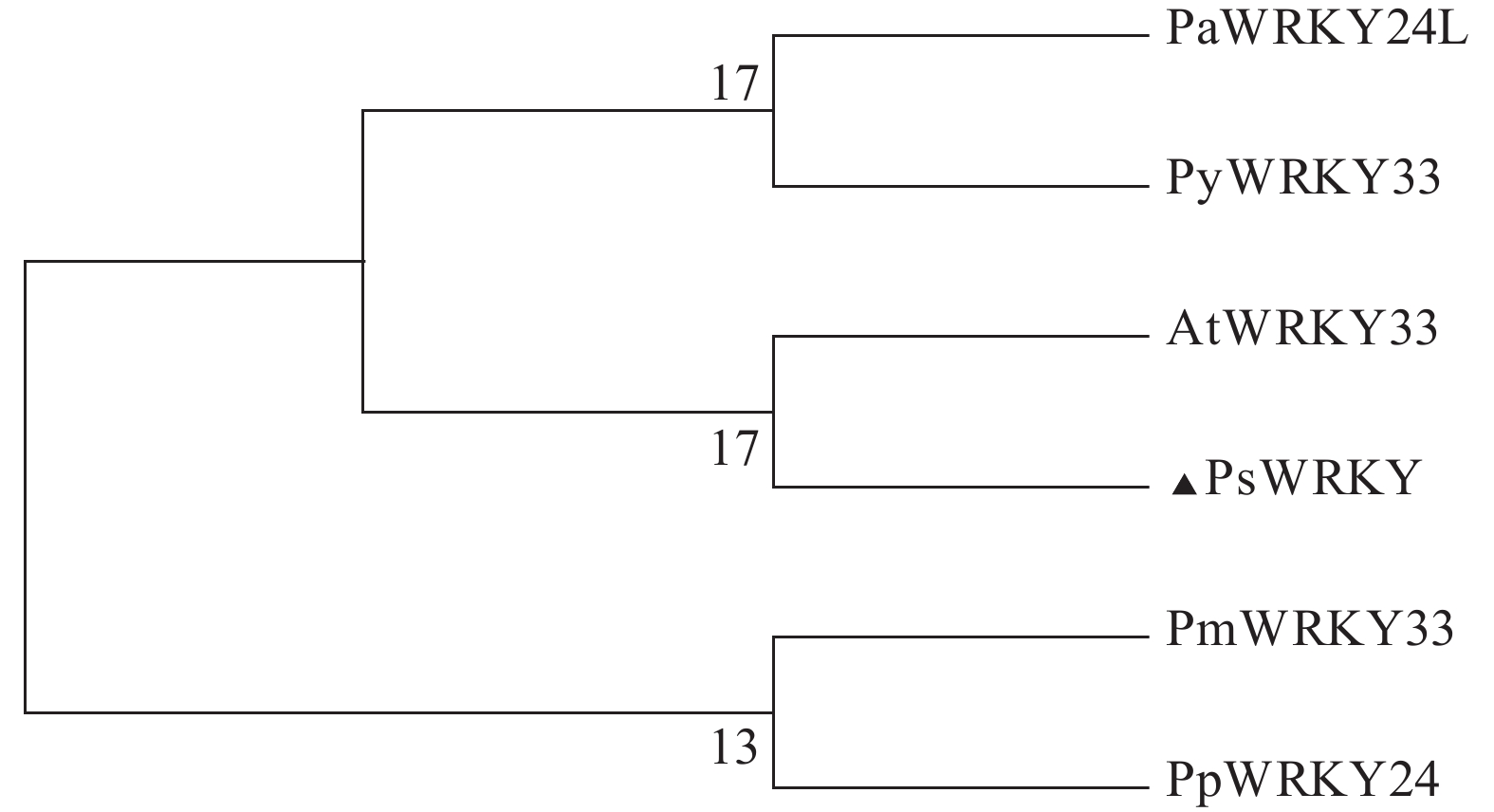
启动子在不同胁迫处理下的表达模式,为进一步深入研究油柰WRKY基因在油柰生长发育或抵御各种逆境胁迫中的作用机制等提供理论依据。 方法 利用MEGA 6.06软件构建PsWRKY33的系统进化树。通过染色体步移技术克隆获得该基因启动子序列,利用PlantCARE数据库分析预测PsWRKY335′端上游启动子区域的顺式作用元件。利用拟南芥浸花法获得转基因植株。通过对不同胁迫处理下的各片段转基因幼苗进行组织化学染色和GUS酶活性测定。 结果 系统进化树分析表明PsWRKY蛋白与拟南芥WRKY33亲缘关系最近,故将该基因命名为PsWRKY33。获得PsWRKY33 基因5′端上游启动子序列长度为1 872 bp,经预测分析发现该启动子区域含有ABRE、ARE、LTR、MYB和W-box等响应不同植物激素等顺式作用元件,基于此构建3个不同长度的缺失片段。对转基因植株不同组织GUS染色发现PsWRKY33 启动子主要表达在叶、花瓣和花序梗上,且随着片段的缺失其表达越低。在不同胁迫处理下,其GUS活性表达程度不同。在低温胁迫条件下,不同长度的PsWRKY33启动子片段均受到不同程度的诱导上调表达;而在SA胁迫下,各片段呈现出不同程度的诱导下调表达。 结论 PsWRKY33基因启动子可能参与调节油柰应答低温胁迫及外源激素SA的响应。 Abstract:Objective Expression of PsWRKY33promoter of Nane (Prunus salicina lindley) under stress was determined for a study on the mechanism of WRKY gene associated with the growth and stress-resistance of the plant. Method Unrooted phylogenetic tree representing relationships among protein sequences of PsWRKY33 was constructed using Nane and other plants through by MEGA 6.06 software. The promoter sequence was cloned by chromosome stepping technique. The cis-acting regulatory elements in the upstream promoter region at the 5′ end of PsWRKY33 were predicted according to the PlantCARE database. Transgenic plants were obtained by using the Arabidopsis immersion method. Histochemical staining and GUS enzyme activity were performed on the transgenic seedlings under different stresses.Result The phylogenetic tree analysis showed that the PsWRKY protein was most closely related to Arabidopsis WRKY33, thus the gene was named PsWRKY33. The 1 872 bp length of 5′ end upstream in PsWRKY33 was obtained, from Nane via genome walking technology. The promoter region contained motifs of hormones (ABRE and ARE) and biotic stresses (LTR, MYB, W-box, and other cis-acting elements). Then, the 3 5′-deletion fragments were constructed. The histochemical staining and GUS activity measurement showed that the full PsWRKY33 promoter fragment mainly expressed in leaves, petals, and peduncles, and its expression decreased with the loss of fragments. Under low temperature, the various fragments of PsWRKY33 promoter were upregulated, but downregulated under SA stress.ConclusionPsWRKY 33 might response to low temperature and exogenous hormone SA stress in Nane.-
Keywords:
- Prunus salicina /
-
PsWRKY33
- promoter /
- histochemical staining /
- stress
-
0. 引言
【研究意义】常规水稻育种方法需要经过多代分离逐渐形成稳定的品种,而花药培养技术具有缩短育种年限,快速稳定育种材料等优点,利用花培技术作为育种辅助手段在创制原始材料、选育新品种、提纯不育系等方面成效显著[1-2]。籼稻和粳稻是水稻的两个亚种,亲缘关系较远,它们在株型、品质和抗逆等方面具有丰富的多样性,通过挖掘两个亚种之间的杂种优势,有助于在抗逆和品质方面进一步提高,同时也是水稻功能基因挖掘的重要基因资源。通过花药培养技术能够快速获得稳定的籼粳育种中间材料,也是构建籼粳交DH遗传群体的有效方法[3-5]。【前人研究进展】一般来说粳稻花药更容易诱导,籼稻材料较难形成愈伤组织,绿苗率低,而籼粳交材料花药培养难度介于二者之间[6-7]。前人的研究发现,供体品种的基因型是花药培养最关键因素,目前认为单核靠边期取样配合低温处理是水稻花培最常规有效的处理方式[6, 8]。同时,培养基类型是目前花药培养效率重要的变化因素[9-10]。施利利等[11]以津稻291/ IRBB60籼粳正反交材料作为研究材料,发现SK3培养基具有最高的愈伤诱导率。吴丹等[7]以5份不同籼粳成分的水稻为试验材料,分别以不同基本培养基(改良M8、SK3、合5、NB)、碳源(蔗糖和麦芽糖)、激素(单一激素、复合激素)为变量,设计16种诱导培养基进行花药培养,发现不同培养基的诱导率和成苗率存在显著差异,以M8为基本培养基,添加蔗糖为碳源,复合激素能获得较高成苗率。何涛等[12]利用21个不同基因型的水稻材料对不同培养条件的花药培养力进行比较,发现基因型对花药培养力影响最大,但是3种培养基在各种基因材料诱导率方面各不相同,没有发现一种较为通用的培养基配方。【本研究切入点】云引是具有广谱稻瘟病抗性的粳稻品种,田间注射接菌和室内离体接菌结果表明其对大多数稻瘟病具有抗性[13],是一个优良的花培供体亲本。然而,相同的水稻材料,在不同培养基成分构成的条件下,如培养基、碳源、植物生长调节剂配比,花药的诱导率和成苗率等都存在显著差异[6-7]。此外,前人的研究往往将愈伤诱导和分化成苗的条件单独进行考虑,缺乏对不同愈伤诱导条件下绿苗成苗率进行研究。【拟解决的关键问题】为了提高云引籼粳杂交后代花药培养效率,本研究拟通过优化培养基成分以及分化愈伤大小等因素,增加花药培养绿苗率以及花培力,为稻瘟病抗性品种云引的挖掘和利用奠定基础。
1. 材料与方法
1.1 材料创制
选用本实验室保存的稻瘟病抗性粳稻品种云引作为母本,与常规籼稻品种粤晶丝苗进行杂交,在田间种植F1代植株,取F1幼穗进行后续试验。
1.2 花药培养流程
当水稻处于孕穗期、叶枕距为5~15 cm时(此时期一般为花粉分化的单核靠边期),剪取穗部,用75%酒精进行表面消毒,并用湿滤纸包裹,置于7℃低温处理10 d;在超净台中将叶鞘剥开,将幼穗取出剪成5 cm长度,置于250 mL三角瓶中,用75%酒精消毒1 min,用无菌水冲洗1遍,用2%次氯酸钠消毒10 min,期间轻轻震荡数次,随后用无菌水将残余的次氯酸钠冲洗干净;用尖头镊子剥开花药,将花药接种于90 mm培养皿中,每个培养皿接种花药100枚,放置于27℃暗培养20~50 d,待愈伤组织长出后,转移到分化培养基上,分化15~30 d,统计绿苗数量。
1.3 培养基配方
诱导培养基:采用常用的NB培养基[14]或M8培养基[15]作为基础培养基,选择麦芽糖或蔗糖作为碳源(质量浓度50 g·L−1),选用复合植物生长调节剂含不同质量浓度2,4-D(0.5~2.0 mg·L−1)、1.0 mg·L−1 KT以及1.0 mg·L−1 NAA,添加3.0 g·L−1植物凝胶,pH=5.85。诱导培养基详见表1。
表 1 培养基成分Table 1. Components in culture media培养基编号
NO. of media基础培养基
Basic media碳源
Carbon source2,4-D质量浓度
Concentration of 2,4-D/(mg·L−1)NS05 NB 蔗糖 0.5 NS10 NB 蔗糖 1.0 NS20 NB 蔗糖 2.0 MS05 M8 蔗糖 0.5 MS10 M8 蔗糖 1.0 MS20 M8 蔗糖 2.0 NM05 NB 麦芽糖 0.5 NM10 NB 麦芽糖 1.0 NM20 NB 麦芽糖 2.0 MM05 M8 麦芽糖 0.5 MM10 M8 麦芽糖 1.0 MM20 M8 麦芽糖 2.0 分化培养基由MS基础培养基的大量元素和微量元素,添加M8有机成分、20 g·L−1蔗糖和20 g·L−1山梨醇、3 mg·L−1 KT和0.5 mg·L−1 NAA、8.8 g·L−1琼脂,pH=5.9。
1.4 数据分析与体系确定
根据花药培养力选择适合于云引籼粳交材料最适合培养基和培养条件,其中出愈率=愈伤数/花药数×100%,绿苗率=绿苗数/愈伤数×100%,得苗率=(绿苗数+白化苗数)/愈伤组织数×100%,花药培养力=出愈率×绿苗率×100%。
2. 结果与分析
2.1 基础培养基对培养力的影响
基础培养基的成分不同,对于愈伤组织的出愈率影响较大,进而影响花药培养力。从表2可以看出,M8培养基无论在出愈率还是在绿苗率上都显著高于NB培养基,其中NB的最高花药培养力仅为0.52%,而M8的最高花药培养力接近2%,花药培养效率相差近4倍,因此后续选择M8作为云引籼粳交的基础培养基。
表 2 不同培养基组合的花培力的比较Table 2. Anther cultivation performances on media of varied formulations培养基
Media基础培养基
Basic media出愈率
Rate of callus formation/%绿苗率
Rate of green seedling/%花药培养力
Anther culture ability/%NS05 NB 2.32 e N/A N/A NS10 NB 5.88 dc 8.64 d 0.51 d NS20 NB 7.34 bc 7.32 d 0.52 d MS05 M8 6.75 bc 19.6 b 1.32 b MS10 M8 13.32 a 14.2 c 1.89 a MS20 M8 16.75 a 11.5 c 1.92 a NM05 NB 1.35 f N/A N/A NM10 NB 1.32 f N/A N/A NM20 NB 3.28 e N/A N/A MM05 M8 3.75 e 24.3 a 0.90 c MM10 M8 6.36 c 21.3 ab 1.34 b MM20 M8 8.23 b 17.5 b 1.40 b 注:①同列数据后含不同字母者在5%水平上差异显著,表3同。②N/A表示样本过少而不被采用。
Note: ① Figures marked with same letter on a same column are not significantly different at 5% level. Same for Table 3. ② N/A indicates sample size is too low to be useful.2.2 不同碳源对培养力的影响
比较了麦芽糖和蔗糖作为碳源对花药培养力的影响(表2)。在M8基础培养基条件下,麦芽糖的绿苗率比蔗糖高,但是蔗糖具有较高的出愈率。综合来看,以蔗糖为碳源具有较高的花培力。此外,本研究同时比较了出愈早期(28 d)和出愈后期(42 d)的愈伤数(表3),结果表明以蔗糖为碳源在出愈早期就能产生较多的愈伤组织,而以麦芽糖为碳源出愈早期的平均出愈率仅相当于出愈后期的50%,因此以蔗糖作为碳源较为合适。
表 3 不同诱导时间的出愈率比较Table 3. Callus induction rates by varied induction time培养基编号
NO. of media28 d出愈率
Rate of callus formation in 28 d/%42 d出愈率
Rate of callus formation in 42 d/%28 d出愈率/
42 d出愈率
Ration of callus formation (28 d versus 42 d) /%MS05 2.52 cd 4.25 d 59.29 MS10 7.25 b 9.35 b 77.54 MS20 11.32 a 14.50 a 78.07 MM05 1.45 d 3.25 d 44.62 MM10 3.22 c 5.35 cd 60.19 MM20 3.35 c 7.35 bc 45.58 2.3 不同2,4-D质量浓度对于绿苗率的影响
在M8基础培养基条件下,随着2,4-D质量浓度增高,出愈率显著上升,而绿苗率下降(表2、3)。以M8+蔗糖为例,添加2 mg·L−1的2,4-D能够获得更高的出愈率,但是绿苗率低于添加1 mg·L−1 2,4-D的组合,因此二者花药培养力相当,相似的情况也出现在M8+麦芽糖体系中(表2)。而在M8+麦芽糖体系中,添加0.5 mg·L−1的2,4-D虽然具有较低的早期出愈率,但1.0 mg·L−1与2.0 mg·L−1的2,4-D基本无差异。因此在花培时,应该综合考虑出愈率和绿苗率,来选择合适的2,4-D质量浓度水平。
2.4 不同愈伤组织大小的绿苗率
为了探讨不同愈伤组织大小对于绿苗率的影响。选取在MM20培养基上,出愈伤组织时间为35~42 d的愈伤组织,待愈伤组织长至0.1、0.2、0.4、0.8 cm时,转移至分化培养基,分化30 d后统计绿苗率(表4),其中0.1 cm大小的愈伤组织,分化得苗率最低,仅为23.2%,而绿苗率约为4.1%,表明愈伤组织过小成苗率显著下降,大部分愈伤组织褐化死亡;0.2 cm大小的愈伤组织,得苗率为63.2%,但是绿苗率高达18.3%;0.4 cm的愈伤组织,得苗率高达85.5%,但是大部分为白化苗,其绿苗率下降为13.3%;0.8 cm的愈伤组织,得苗率为83.6%,但其绿苗率仅为5.35%。因此,当愈伤长至0.2 cm时进行分化是最佳时机,愈伤组织在培养基上生长时间不宜过长,否则分化力显著下降。
表 4 不同愈伤组织大小进行分化成苗得苗率的比较Table 4. Seedling yields after differentiation using calluses of different sizes(单位/%) 组别
Groups愈伤组织大小 Callus Size 0.1 cm 0.2 cm 0.4 cm 0.8 cm 全部 Total seedlings 23.2 63.2 85.5 83.6 绿苗 Green seedlings 4.1 18.3 13.3 5.35 3. 讨论与结论
综合上述的试验结果,云引籼粳交材料的花药培养条件为:M8基础培养基,添加50 g·L−1蔗糖作为碳源和1~2 mg·L−1 2,4-D,当花药愈伤组织长至0.2 cm时转移至分化培养基分化,建立了较为稳定的云引籼粳材料花培体系。
水稻组织培养受基因型影响较大,无论是胚诱导还是花药诱导,粳稻培养力都比籼稻高。而培养条件对于花药培养力影响较大。从本研究来看,对花药培养力的影响,基础培养基成分>碳源>2,4-D 质量浓度,而绿苗率随植物生长调节剂浓度增加而降低。
同时,愈伤组织分化大小也会影响绿苗率以及得苗率。虽然水稻花药培养技术已较为成熟,但是选择合适的培养基能够显著提高花药培养力,特别是能够提高绿苗率。粳稻出愈率普遍较高,本课题组在前期对其他粳稻品种进行花药诱导时发现,绿苗分化率并不显著高于籼稻品种。前人的研究也发现,籼粳成分高低与绿苗分化率无显著相关性[7, 12]。因此如何提高绿苗率是花药培养优化的重要方向,应当特别注意:虽然高质量浓度的2,4-D能够提高出愈率,且在高质量浓度2,4-D条件下愈伤组织生长虽然不受影响,但是绿苗率显著下降。今后或许可以通过分步培养,即前期高质量浓度2,4-D诱导,后期转至低质量浓度2,4-D生长的培养方法来提高绿苗率,进而提高花药培养的效率。
-
图 2 PsWRKY33
基因启动子序列 注:-60表示ATG上游60个核苷酸。红色粗体字母加框表示预测的顺式作用元件序列。pPsWRKY33、P1、P2、P3加箭头加黑色下划线表示为4个片段(−1 872、−852、−357和−271 bp)的特异性引物,箭头表示它们的方向。
Figure 2. Cis-elements of PsWRKY33 promoter
Note: -60 represented the 60 nucleotides upstream of the ATG. Bold red letters with boxes indicated the predicted sequence of cis-acting elements. pPsWRKY33, P1, P2 and P3 add arrows and black underlines represented the specific primers of four fragments (−1 872, −852, −357 and −271 bp), the arrows indicated their direction.
图 3 PsWRKY33
启动子各片段的GUS载体 注:不同启动子区域的数目表示PsWRKY33翻译起始ATG密码子上游的核苷酸位置。attR1和attR2限制性位点:将pDMC163载体中的CaMV35S启动子替换为PsWRKY33启动子片段。LB和RB:代表T-DNA的左右边缘;Hyg:潮霉素标记基因;GUSA:GUS基因编码区域。黄色的条形图代表PsWRKY33::GUS、PsWRKY33-P1::GUS、PsWRKY33-P2::GUS和PsWRKY33-P3::GUS结构的-1 872 bp、-852 bp、-357 bp和-271 bp PsWRKY33启动子片段。
Figure 3. Schematic diagram of GUS vector of individual PsWRKY33 promoter fragments
Note: The number of different promoter regions represents the nucleotide position upstream of the PsWRKY33 translation initiation ATG codon. The CaMV35S promoter in pDMC163 vector was replaced by PsWRKY33 promoter fragment by attR1 and attR2 restriction sites. LB and RB represent the left and right edges of T-DNA; Hyg, hygromycin B marker gene; GUS is the region that codes for the GUS gene. The yellow bars represent the -1 872 bp, -852 bp, -357 bp and -271 bp PsWRKY33 promoter fragments of PsWRKY33::GUS, PsWRKY33-P1::GUS, PsWRKY33-P2::GUS and PsWRKY33-P3::GUS, respectively.
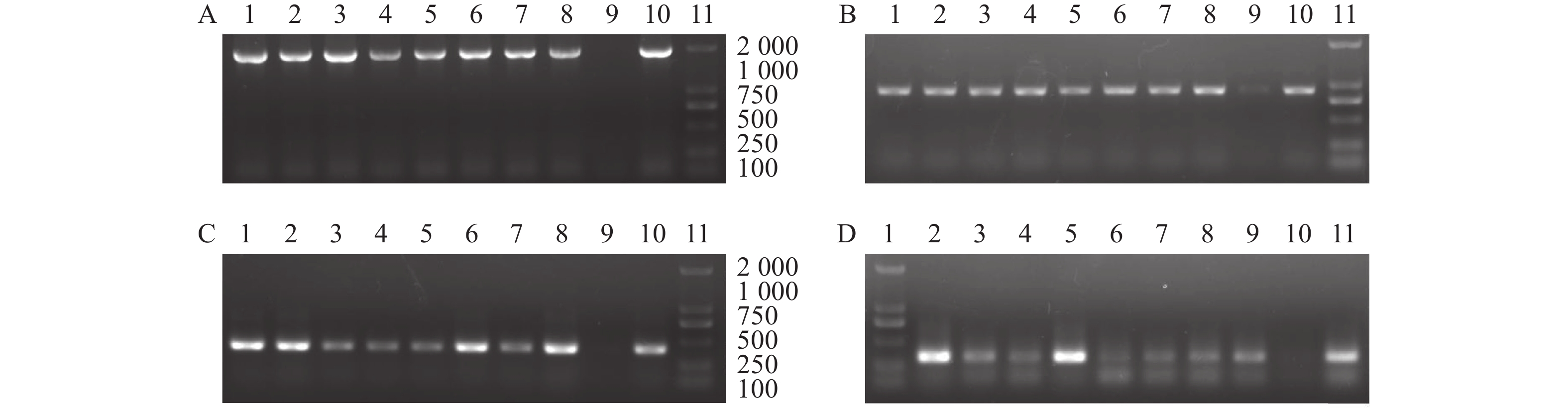
图 4 PCR检测T2代转基因植株
注:A~D分别为pPsWRKY33、pPsWRKY33-P1、pPsWRKY33-P2和pPsWRKY33-P3转基因植株的PCR检测结果。1~8分为pPsWRKY33、pPsWRKY33-P1、pPsWRKY33-P2和pPsWRKY33-P3 8个转基因植株;9为质粒;10为野生型;11为DL 2 000 DNA maker。
Figure 4. Detectioned of transgenic Arabidopsis T2 plants by PCR
Note: A-D: PCR analysis on pPsWRKY33, pPsWRKY33-P1, pPsWRKY33-P2, and pPsWRKY33-P3 transgenic Arabidopsis, respectively; 1-8: 8 strains of pPsWRKY33, pPsWRKY33-P1, pPsWRKY33-P2, and pPsWRKY33-P3 transgenic Arabidopsis, respectively; 9: plasmid; 10: wild type; 11: DNA maker DL 2 000.
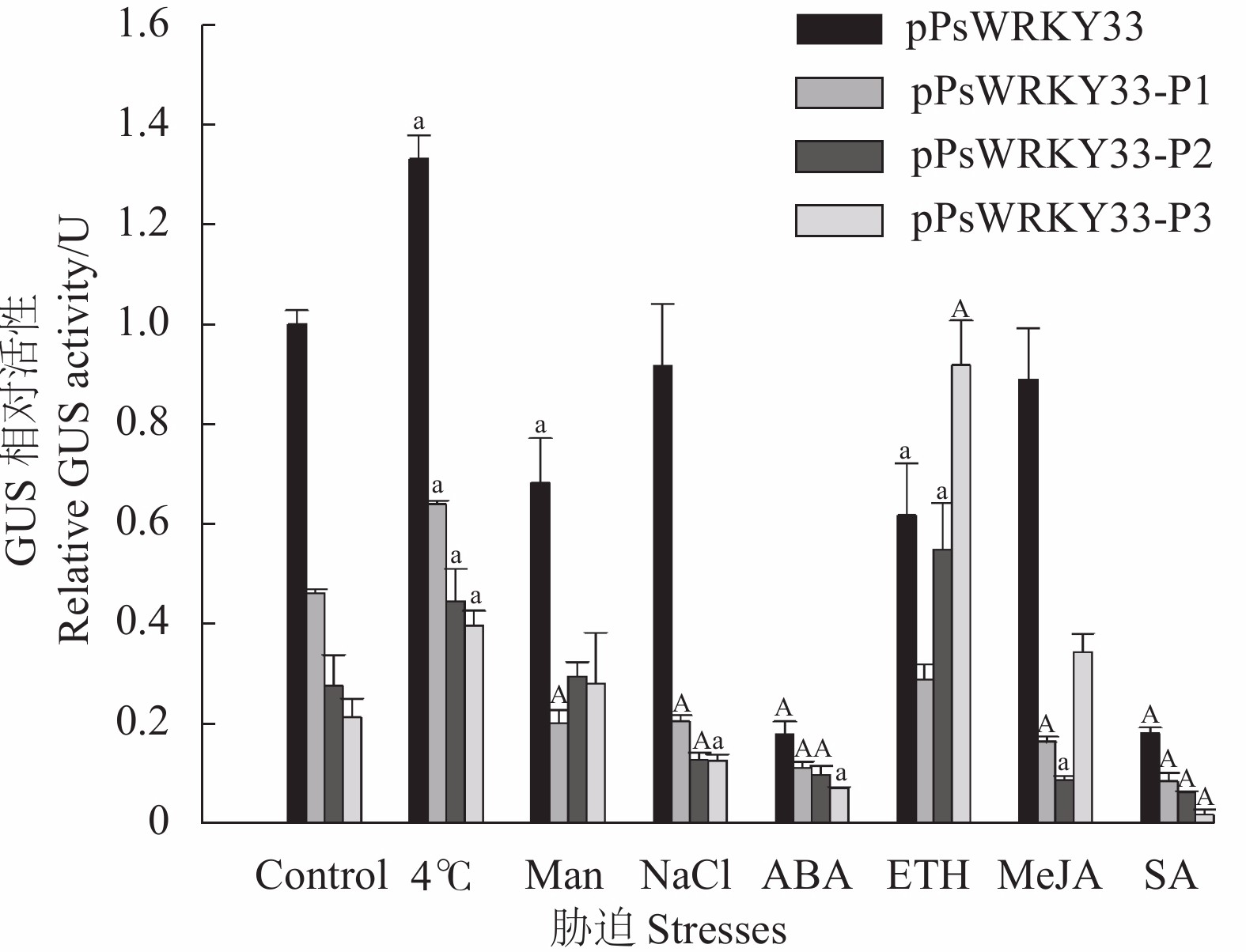
图 7 转基因拟南芥在不同胁迫处理下的GUS相对活性
注:图中不同大、小写字母表示同一处理达极显著(P<0.01)、显著(P<0.05)差异。
Figure 7. GUS relative activity in transgenic Arabidopsis under different stresses
Note: Different uppercase and lowercase letters indicate highly significant (P<0.01) and significant (P<0.05) differences in the same treatment .
表 1 引物序列
Table 1 Sequences of primers
引物名称 Primer name 引物序列(5′-3′) Sequence of primers(5′-3′) PsWRKY33-GSPA GCGAAGTAAGAAGAGGGAGAAACGGAG PsWRKY33-GSPB TGTTGGTGCTGGGGTCTTCATCTG PsWRKY33-GSPC CGAGAAAGTGAAGGTTGGGTGTGAA pPsWRKY33-F GGGGACAAGTTTGTACAAAAAAGCAGGCTTCGAAGTTCTGCCTTGTTTGCT pPsWRKY33-P1-F GGGGACAAGTTTGTACAAAAAAGCAGGCTTCTGATCGAATTTTTTTTTCATA pPsWRKY33-P2-F GGGGACAAGTTTGTACAAAAAAGCAGGCTTCATTATATTGGTTTACATACA pPsWRKY33-P3-F GGGGACAAGTTTGTACAAAAAAGCAGGCTTCAAACCCACGAGCTTTGACCA pPsWRKY33-R GGGGACCACTTTGTACAAGAAAGCTGGGTCAAAGGGGGGGAGAGAGAGAG -
[1] 贾丽. 福建省柰、李、桃种质资源的ISSR分子分析[D]. 福州: 福建农林大学, 2009. JIA L. ISSR analysis of germplasm resources of prunus[D]. Fuzhou: Fujian Agriculture and Forestry University, 2009. (in Chinese)
[2] 王连延. 福建省古田县果树的霜冻灾害 [J]. 落叶果树, 2012, 44(3):48−49. DOI: 10.3969/j.issn.1002-2910.2012.03.025 WANG L Y. Frost disaster of fruit trees in Gutian County, Fujian Province [J]. Deciduous Fruits, 2012, 44(3): 48−49.(in Chinese) DOI: 10.3969/j.issn.1002-2910.2012.03.025
[3] ZHU J K. Abiotic stress signaling and responses in plants [J]. Cell, 2016, 167(2): 313−324. DOI: 10.1016/j.cell.2016.08.029
[4] 钟新华. 油柰低产成因及改造技术调查 [J]. 福建果树, 1999(4):24−25. ZHONG X H. Investigation of genesis and transformation technology of lower yield in Oil Nai [J]. Fujian Fruits, 1999(4): 24−25.(in Chinese)
[5] BANERJEE A, ROYCHOUDHURY A. WRKY proteins: Signaling and regulation of expression during abiotic stress responses [J]. The Scientific World Journal, 2015, 2015: 807560.
[6] PHUKAN U J, JEENA G S, SHUKLA R K. WRKY transcription factors: Molecular regulation and stress responses in plants [J]. Frontiers in Plant Science, 2016, 7: 760.
[7] JIANG J J, MA S H, YE N H, et al. WRKY transcription factors in plant responses to stresses [J]. Journal of Integrative Plant Biology, 2017, 59(2): 86−101. DOI: 10.1111/jipb.12513
[8] 相立, 赵蕾, 王玫, 等. 苹果MdWRKY74的克隆和功能分析[J/OL]. 园艺学报, 2021: 1-11 [2021-05-18]. http://kns.cnki.net/kcms/detail/11.1924.s.20210518.1046.002.html. XIANG L, ZHAO L, WANG M, et al. Cloning and functional analysis of MdWRKY74 in apple [J/OL]. Acta Horticulturae Sinica, 2021: 1-11. [2021-05-18]. http://kns.cnki.net/kcms/detail/11.1924.s.20210518.1046.002.html. (in Chinese)
[9] LI S J, FU Q T, CHEN L G, et al. Arabidopsis thaliana WRKY25, WRKY26, and WRKY33 coordinate induction of plant thermotolerance [J]. Planta, 2011, 233(6): 1237−1252. DOI: 10.1007/s00425-011-1375-2
[10] HE G H, XU J Y, WANG Y X, et al. Drought-responsive WRKY transcription factor genes TaWRKY1 and TaWRKY33 from wheat confer drought and/or heat resistance in Arabidopsis [J]. BMC Plant Biology, 2016, 16(1): 116. DOI: 10.1186/s12870-016-0806-4
[11] LI Z, LIANG F P, ZHANG T B, et al. Enhanced tolerance to drought stress resulting from Caragana korshinskii CkWRKY33 in transgenic Arabidopsis thaliana [J]. BMC Genomic Data, 2021, 22(1): 11. DOI: 10.1186/s12863-021-00965-4
[12] 张锦锦. MsWRKY33转录因子调控紫花苜蓿耐盐性的功能研究[D]. 北京: 中国农业科学院, 2021. ZHANG J J. The study on the function of MsWRKY33 transcription factor in regulating salt tolerance of Medicago sativa. L[D]. Beijing: Chinese Academy of Agricultural Sciences, 2021. (in Chinese)
[13] SUN X M, ZHANG L L, WONG D C J, et al. The ethylene response factor VaERF092 from Amur grape regulates the transcription factor VaWRKY33, improving cold tolerance [J]. The Plant Journal:for Cell and Molecular Biology, 2019, 99(5): 988−1002.
[14] 陈永萍, 林金辉, 刘志钦, 等. 油柰PsWRKY22基因启动子的克隆与功能鉴定 [J]. 分子植物育种, 2021, 19(9):2837−2845. CHEN Y P, LIN J H, LIU Z Q, et al. Cloning and functional identification of promoter of PsWRKY22 gene in Nai [J]. Molecular Plant Breeding, 2021, 19(9): 2837−2845.(in Chinese)
[15] 蒋琦, 高广春, 吴霁蓂, 等. WRKY转录因子调控植物次生代谢的研究进展[J/OL]. 分子植物育种, 2021: 1-8 [2021-12-03]. http://kns.cnki.net/kcms/detail/46.1068.S.20211201.1439.006.html. JIANG Q, GAO G C, WU J M, et al. Research progress of WRKY transcription factor regulating plant secondary metabolism [J/OL]. Molecular Plant Breeding, 2021: 1-8 [2021-12-03]. http://kns.cnki.net/kcms/detail/46.1068.S.20211201.1439.006.html. (in Chinese)
[16] 张凡, 尹俊龙, 郭瑛琪, 等. WRKY转录因子的研究进展 [J]. 生物技术通报, 2018, 34(1):40−48. ZHANG F, YIN J L, GUO Y Q, et al. Research advances on WRKY transcription factors [J]. Biotechnology Bulletin, 2018, 34(1): 40−48.(in Chinese)
[17] 王雪, 王盛昊, 于冰. 转录因子和启动子互作分析技术及其在植物应答逆境胁迫中的研究进展 [J]. 中国农学通报, 2021, 37(33):112−119. DOI: 10.11924/j.issn.1000-6850.casb2021-0563 WANG X, WANG S H, YU B. Interaction analysis of transcription factors and promoters and its application in response of plants to stress [J]. Chinese Agricultural Science Bulletin, 2021, 37(33): 112−119.(in Chinese) DOI: 10.11924/j.issn.1000-6850.casb2021-0563
[18] 杨晓娜, 赵昶灵, 李云, 等. 启动子序列克隆和功能分析方法的研究进展 [J]. 云南农业大学学报(自然科学版), 2010, 25(2):283−290. YANG X N, ZHAO C L, LI Y, et al. Research advances in the methods of cloning and function-analyzing of promoters [J]. Journal of Yunnan Agricultural University (Natural Science), 2010, 25(2): 283−290.(in Chinese)
[19] 蔡韡韡. 水杨酸诱导下(木奈)抗性相关基因的分离与表达研究[D]. 福州: 福建农林大学, 2016. CAI W W. Isolation and expression of resistance-associated genes under induction of salicylic acid in (Prunus salicina lindli. var cordata J. Y. Zhang et al.)[D]. Fuzhou: Fujian Agriculture and Forestry University, 2016. (in Chinese)
[20] 高永峰, 杨丰铭, 李琴中, 等. 番茄SlWRKY31基因启动子的克隆与逆境应答模式分析 [J]. 西北植物学报, 2018, 38(12):2155−2164. GAO Y F, YANG F M, LI Q Z, et al. Cloning and analysis of stress response pattern of SlWRKY31 gene promoter from tomato [J]. Acta Botanica Boreali-Occidentalia Sinica, 2018, 38(12): 2155−2164.(in Chinese)
[21] 刘志钦, 杨晟, 蔡金森, 等. 辣椒CaWRKY5启动子的分离及其调控元件分析 [J]. 应用与环境生物学报, 2013, 19(3):389−394. DOI: 10.3724/SP.J.1145.2013.00389 LIU Z Q, YANG S, CAI J S, et al. Isolation and Cis-acting analysis of the CaWRKY5 promoter in pepper [J]. Chinese Journal of Applied and Environmental Biology, 2013, 19(3): 389−394.(in Chinese) DOI: 10.3724/SP.J.1145.2013.00389
[22] 杨鹏芳, 段国琴, 胡晓炜, 等. 高等植物启动子研究概述 [J]. 分子植物育种, 2018, 16(5):1482−1493. YANG P F, DUAN G Q, HU X W, et al. Overview of higher plant promoters research [J]. Molecular Plant Breeding, 2018, 16(5): 1482−1493.(in Chinese)
[23] JUNG H W, KIM K D, HWANG B K. Identification of pathogen-responsive regions in the promoter of a pepper lipid transfer protein gene (CALTPI) and the enhanced resistance of the CALTPI transgenic Arabidopsis against pathogen and environmental stresses [J]. Planta, 2005, 221(3): 361−373. DOI: 10.1007/s00425-004-1461-9
[24] RUSHTON P J, REINSTÄDLER A, LIPKA V, et al. Synthetic plant promoters containing defined regulatory elements provide novel insights into pathogen- and wound-induced signaling [J]. The Plant Cell, 2002, 14(4): 749−762. DOI: 10.1105/tpc.010412



 下载:
下载: